Methods
All three methods share the same interface and return type. Switch methods by passing a different algorithm struct as the first argument to disaggregate.
Comparison
Benchmarks: 20-year span, output_period=Week(1), 8 threads (Julia 1.12). Time = minimum over 2–3 runs. Memory = dominant working-set matrix (n × matrix_width × 8 B).
| Method | Pros | Cons | n = 10k | n = 100k | n = 1M |
|---|---|---|---|---|---|
Spline | No kernel required; optional tension suppresses oscillation near sparse gaps | Design matrix O(n × n_knots); can oscillate without tension | 12 ms<br>19 MB | 103 ms<br>192 MB | 807 ms<br>1.9 GB |
Sinusoid | Analytical integrals (no quadrature); interpretable parameters (amplitude, phase, trend, anomalies); lowest peak memory | Assumes annual periodicity; poor fit for non-sinusoidal signals | 61 ms<br>2 MB | 133 ms<br>19 MB | 2.4 s<br>192 MB |
GP | Arbitrary KernelFunctions.jl kernels; most flexible | O(n·m·q + m³) Cholesky — memory-limited above n ≈ 50 000 at weekly output | 2.0 s<br>195 MB | 13.3 s<br>1.9 GB | —<br>(>8 GB) |
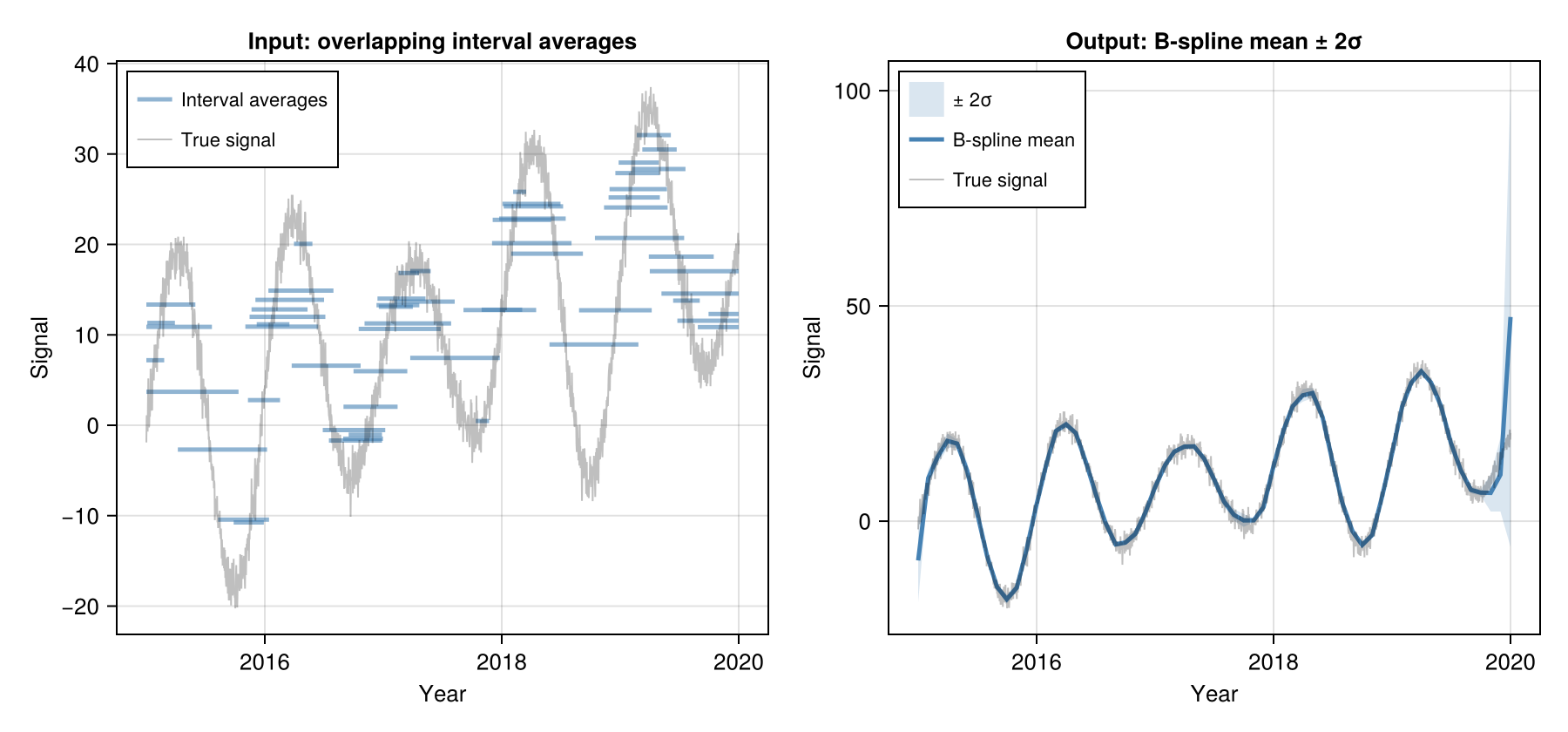
B-spline (Spline)
Fits a smooth curve whose running averages match the observations. A regularisation parameter controls smoothness; an optional tension penalty suppresses oscillation near sparse regions.
result = disaggregate(Spline(
smoothness = 1e-3, # larger = smoother
tension = 0.0, # > 0 suppresses oscillation
penalty_order = 3, # order of difference penalty
), y, t1, t2; loss_norm = L2DistLoss())Uncertainty: Spatially-varying sandwich std — lower where observations are dense, higher where they are sparse.
Tension-spline (Spline with tension > 0)
Adding tension stiffens the curve in data-sparse regions — think of pulling the spline taut like a guitar string. It suppresses oscillation while preserving fidelity where observations are dense.
result = disaggregate(Spline(
smoothness = 1e-3,
tension = 25.0, # 0.5–1 moderate; 5–10 near piecewise-linear; >20 strongly stiffened
), y, t1, t2)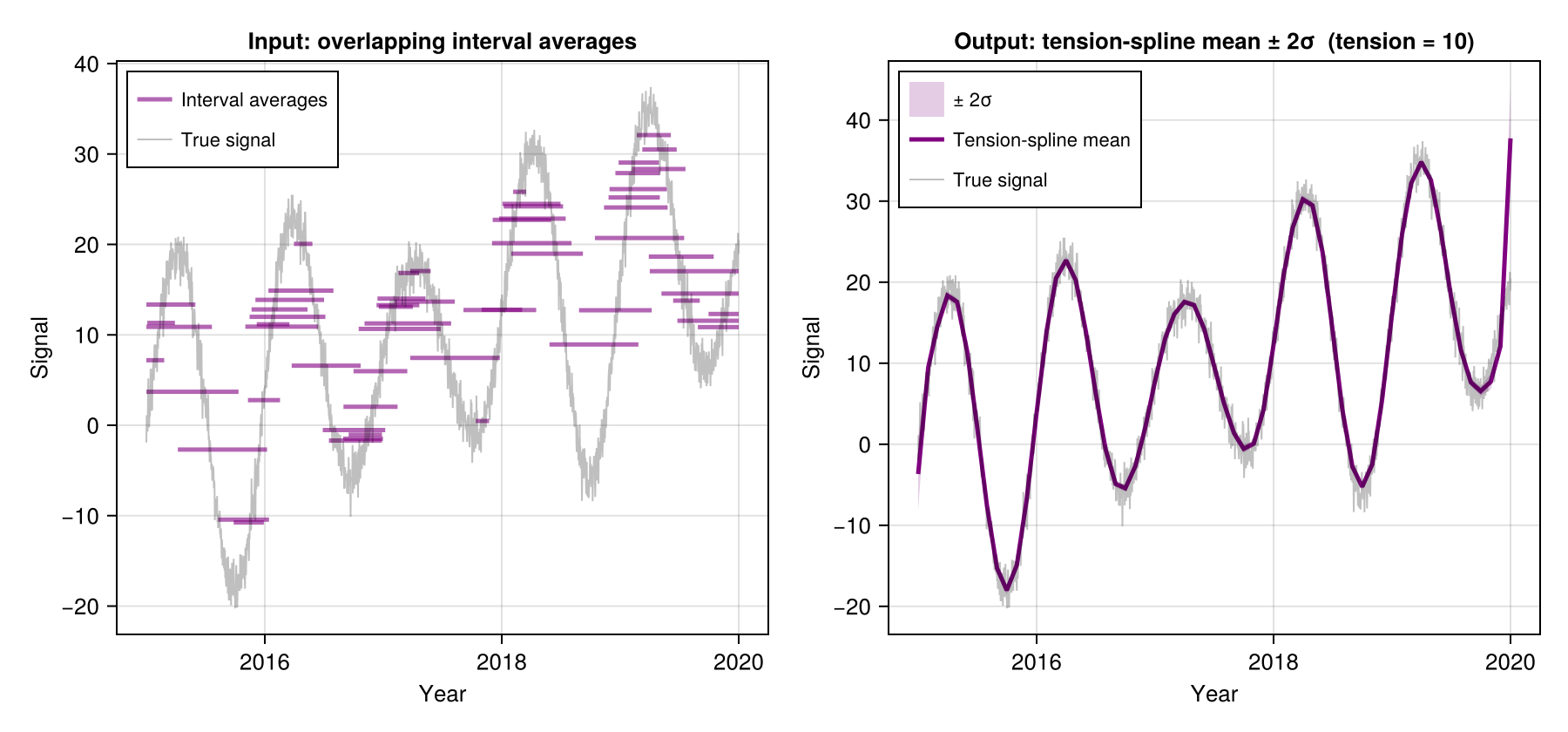
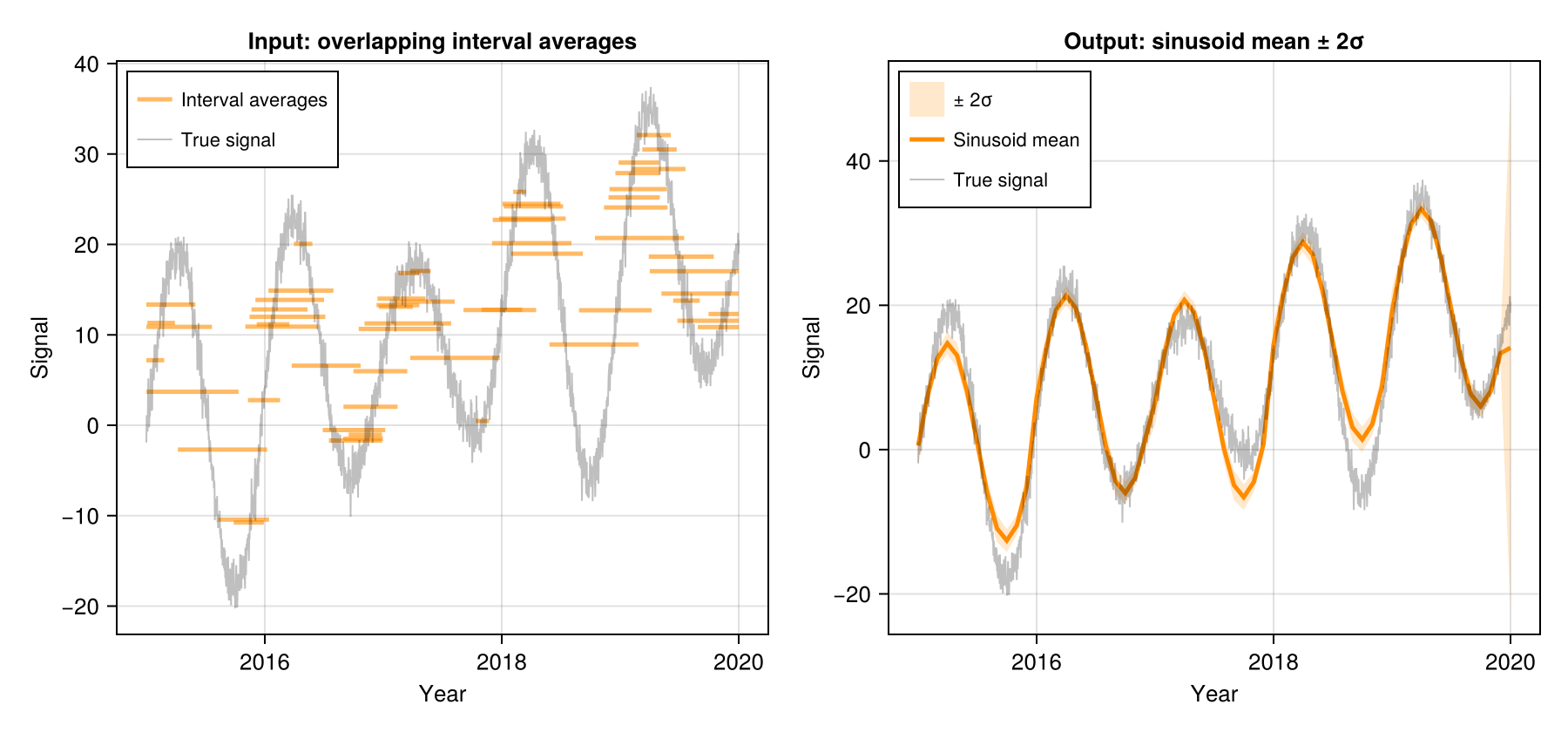
Sinusoid (Sinusoid)
Fits the parametric model:
x(t) = μ + β·(t − t̄) + γ(year) + A·sin(2πt) + B·cos(2πt)where μ is the mean, β is a linear trend, γ(year) is a per-year anomaly, and A, B are annual seasonal amplitudes. All integrals are solved analytically — making this the fastest method.
result = disaggregate(Sinusoid(
smoothness_interannual = 1e-2, # ridge penalty on year-to-year anomalies
), y, t1, t2)
# Fitted parameters
using DimensionalData: metadata
md = metadata(result)
md[:amplitude] # seasonal amplitude √(A²+B²)
md[:phase] # peak time within year (fraction)
md[:trend] # linear trend (units/year)
md[:interannual] # Dict{Int,Float64} of per-year anomaliesUncertainty: Spatially-varying sandwich std — lower where observations are dense, higher where they are sparse.
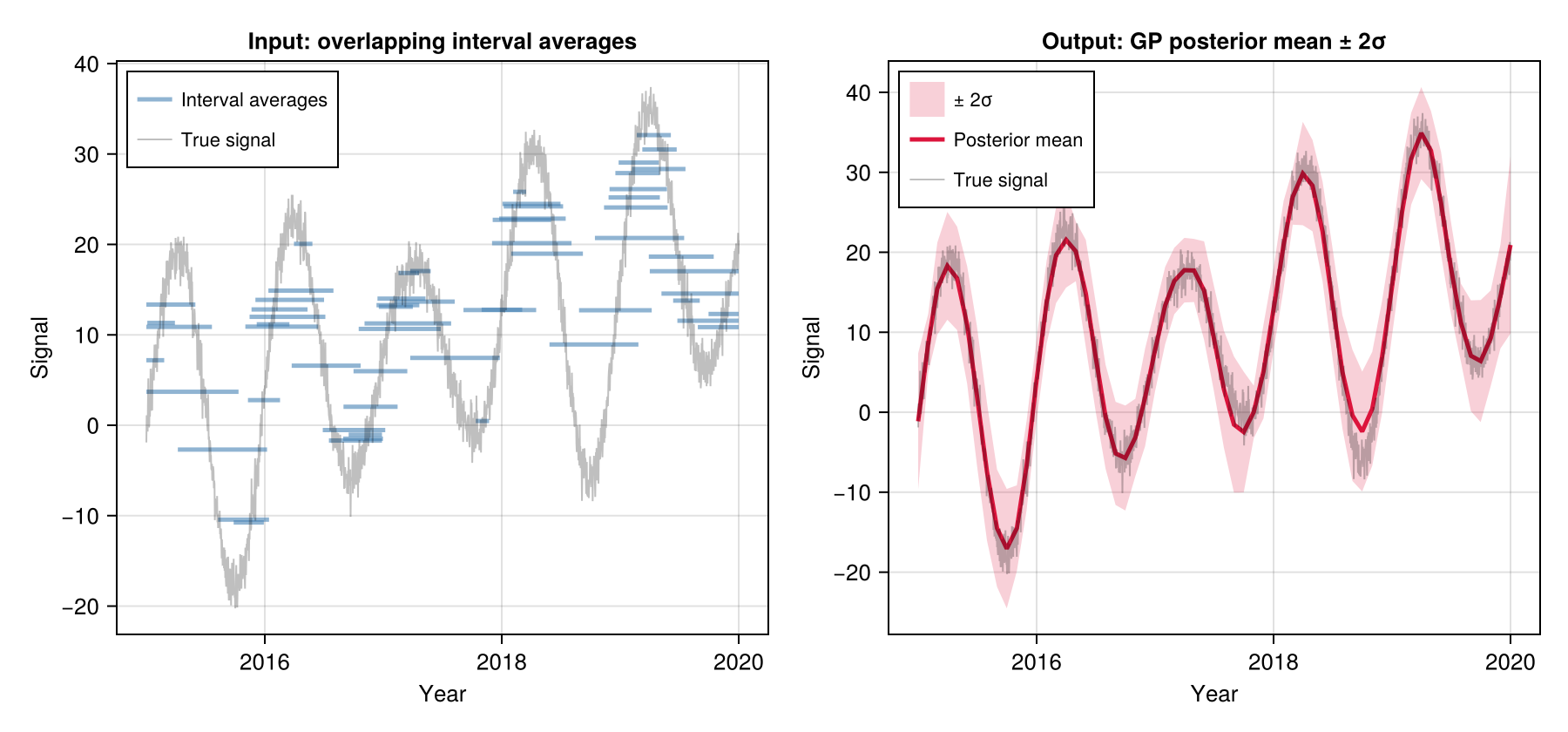
Gaussian Process (GP)
Models the signal as a Gaussian Process — a flexible probabilistic model encoding correlations through time. A sparse approximation keeps computation fast even for long records. Specify the correlation structure via a KernelFunctions.jl kernel.
using KernelFunctions
k = 15.0^2 * PeriodicKernel(r=[0.5]) * with_lengthscale(Matern52Kernel(), 3.0) +
5.0^2 * with_lengthscale(Matern52Kernel(), 2.0)
result = disaggregate(GP(
kernel = k,
obs_noise = 4.0, # initial noise variance σ² (auto-calibrated by default)
n_quad = 5, # Gauss-Legendre quadrature points per interval
calibrate_noise = true, # iteratively adjust obs_noise to match residual RMS
), y, t1, t2)obs_noise calibration: When calibrate_noise=true (default), the method iteratively adjusts obs_noise to the value where the weighted RMS of interval-average residuals equals sqrt(obs_noise). This is an empirical Bayes fixed-point that makes the posterior mean insensitive to the initial obs_noise value — useful when obs_noise is hard to set a priori (e.g. when data units or scale are unknown). Set calibrate_noise=false to use obs_noise exactly as specified.
Uncertainty: Spatially-varying sandwich std — lower where observations are dense, higher where they are sparse.
Robust Loss Functions
TemporalDisaggregations.jl uses LossFunctions.jl for its robust loss implementations. Pass any DistanceLoss type:
L2DistLoss()— Least squares (standard, no IRLS)L1DistLoss()— Absolute deviation (robust to outliers via IRLS)HuberLoss(δ)— Hybrid loss (L2 for small residuals, L1 for large residuals)
Example with custom Huber threshold:
using LossFunctions
result = disaggregate(Spline(), y, t1, t2; loss_norm=HuberLoss(2.0))Under the hood, IRLS weights are computed as w = 1 / (|∂L/∂r| + ε) where ∂L/∂r is provided by LossFunctions.jl's deriv() function.
Uncertainty
All three methods return the same type of std — a spatially-varying sandwich standard deviation:
std(t*) = σ̂ · sqrt(q(t*))where σ̂ is the weighted residual RMS of predicted vs. observed interval averages (sqrt(Σ wᵢ rᵢ² / Σ wᵢ), with rᵢ = yᵢ − ŷᵢ) and q(t*) is a dimensionless coverage factor derived from the method's hat vector at time t*:
Dense observation coverage →
q(t*) < 1→std(t*) < σ̂Sparse observation coverage →
q(t*) > 1→std(t*) > σ̂
This makes std comparable across methods and automatically reflects the temporal distribution of the input observations. When using robust losses (L1DistLoss(), HuberLoss(δ)), both σ̂ and q(t*) are computed from the final IRLS solution.